Supervised Variational Autoencoder (VAE) regression for high‑dimensional predictors (e.g., VIS–NIR–SWIR soil spectroscopy), implemented in Python TensorFlow/Keras and exposed in R via reticulate.
The README is also the main reproducible case study, mirroring the vignette (vignettes/soilVAE-workflow.Rmd) so a reader can understand the why, how, and performance without opening additional files.
What soilVAE does
Given spectra and a continuous soil property , soilVAE learns:
- an encoder mapping spectra to a latent embedding
- a decoder reconstructing spectra
- a supervised head predicting the property
Installation
CRAN (once accepted)
install.packages("soilVAE")Python / TensorFlow setup that does not surprise the user
Because deep learning depends on external Python libraries, this README uses a defensive pattern:
- detect whether Python + TF/Keras are available
- if not, show exactly how to create a conda env using conda-forge only
- run the VAE only when the environment is ready
Important:
reticulate“locks” the Python used per R session. If you change env variables oruse_*()calls, restart R.
Option A (recommended): conda env (conda-forge only)
library(reticulate)
# Make sure reticulate isn't forced to a missing python
Sys.unsetenv("RETICULATE_PYTHON")
# Create env (if needed)
if (!"soilvae-tf" %in% conda_list()$name) {
conda_create("soilvae-tf", python_version = "3.11")
}
# Install core deps from conda-forge
conda_install("soilvae-tf", packages = c("pip", "numpy"), channel = "conda-forge")
# Install TF/Keras via pip inside the env
py_install(c("tensorflow>=2.13", "keras>=3"), pip = TRUE, envname = "soilvae-tf")Now, in the same R session:
library(soilVAE)
soilVAE::vae_configure(conda = "soilvae-tf")
reticulate::py_config()Option B: point to an existing Python executable
library(soilVAE)
soilVAE::vae_configure(python = "C:/path/to/python.exe")Reproducible case study (spectra -> pre-processing -> PLS baseline -> soilVAE)
This follows the workflow style commonly used in soil spectral inference tutorials (e.g., Wadoux et al., 2021) (Wadoux et al. 2021), with a direct comparison between a PLS baseline and soilVAE.
Packages
set.seed(19101991)
pkgs <- c("prospectr", "pls", "reticulate")
for (p in pkgs) if (!requireNamespace(p, quietly = TRUE)) install.packages(p)
library(prospectr)
library(pls)
library(reticulate)
if (!requireNamespace("soilVAE", quietly = TRUE)) {
stop("soilVAE is not installed. Install it with remotes::install_github('HugoMachadoRodrigues/soilVAE').")
}
library(soilVAE)
# Defensive: detect Python + TF/Keras early, so the README can render everywhere.
has_py <- reticulate::py_available(initialize = FALSE)
has_tf <- FALSE
if (has_py) {
try(reticulate::py_config(), silent = TRUE)
has_tf <- reticulate::py_module_available("tensorflow") &&
reticulate::py_module_available("keras")
}Data
This example assumes you ship datsoilspc inside the package (data/datsoilspc.rda).
This dataset is provided and described at Geeves et al. (1995)
Geeves, G. W. (Guy W.) & New South Wales. Department of Conservation and Land Management & CSIRO. Division of Soils. (1995). The physical, chemical and morphological properties of soils in the wheat-belt of southern N.S.W. and northern Victoria / G.W. Geeves … [et al.]. Glen Osmond, S. Aust. : CSIRO Division of Soils
data("datsoilspc", package = "soilVAE")
str(datsoilspc)
#> 'data.frame': 391 obs. of 5 variables:
#> $ clay : num 49 7 56 14 53 24 9 18 33 27 ...
#> $ silt : num 10 24 17 19 7 21 9 20 13 19 ...
#> $ sand : num 42 69 27 67 40 55 83 61 54 55 ...
#> $ TotalCarbon: num 0.15 0.12 0.17 1.06 0.69 2.76 0.66 1.36 0.19 0.16 ...
#> $ spc : num [1:391, 1:2151] 0.0898 0.1677 0.0778 0.0958 0.0359 ...
#> ..- attr(*, "dimnames")=List of 2
#> .. ..$ : NULL
#> .. ..$ : chr [1:2151] "350" "351" "352" "353" ...
#> - attr(*, "na.action")= 'omit' Named int 392
#> ..- attr(*, "names")= chr "63"Expected structure:
-
datsoilspc$spc: matrix of reflectance spectra (rows = samples; cols = wavelengths) -
datsoilspc$TotalCarbon: numeric target (example property)
Utility: evaluation metrics (base R)
We replicate typical “quantitative” metrics used in soil spectroscopy:
RMSE, MAE, R², bias (ME), RPIQ, and RPD.
eval_quant <- function(y, yhat) {
y <- as.numeric(y)
yhat <- as.numeric(yhat)
ok <- is.finite(y) & is.finite(yhat)
y <- y[ok]
yhat <- yhat[ok]
if (length(y) < 3) {
return(list(
n = length(y),
ME = NA_real_, MAE = NA_real_, RMSE = NA_real_,
R2 = NA_real_, RPIQ = NA_real_, RPD = NA_real_
))
}
err <- yhat - y
me <- mean(err)
mae <- mean(abs(err))
rmse <- sqrt(mean(err^2))
ss_res <- sum((y - yhat)^2)
ss_tot <- sum((y - mean(y))^2)
r2 <- if (ss_tot == 0) NA_real_ else 1 - ss_res / ss_tot
rpiq <- stats::IQR(y) / rmse
rpd <- stats::sd(y) / rmse
list(
n = length(y),
ME = me,
MAE = mae,
RMSE = rmse,
R2 = r2,
RPIQ = rpiq,
RPD = rpd
)
}
as_df_metrics <- function(x) {
data.frame(
n = x$n,
ME = round(x$ME, 2),
MAE = round(x$MAE, 2),
RMSE = round(x$RMSE, 2),
R2 = round(x$R2, 2),
RPIQ = round(x$RPIQ, 2),
RPD = round(x$RPD, 2),
stringsAsFactors = FALSE
)
}Plot reflectance spectra
matplot(
x = as.numeric(colnames(datsoilspc$spc)),
y = t(as.matrix(datsoilspc$spc)),
xlab = "Wavelength / nm",
ylab = "Reflectance",
ylim = c(0, 1),
type = "l",
lty = 1,
col = rgb(0.5, 0.5, 0.5, alpha = 0.3)
)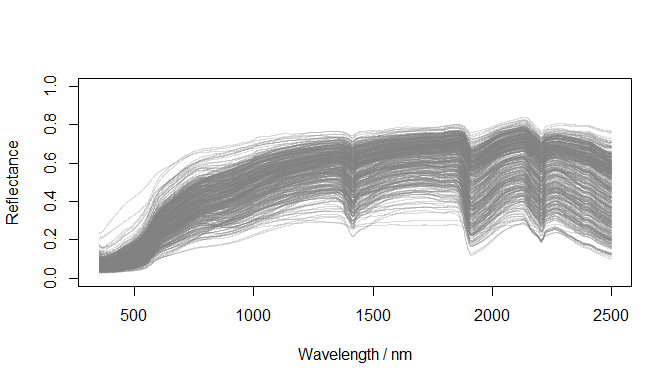
Raw reflectance spectra (datsoilspc$spc)
Convert reflectance to absorbance
datsoilspc$spcA <- log(1 / as.matrix(datsoilspc$spc))
matplot(
x = as.numeric(colnames(datsoilspc$spcA)),
y = t(datsoilspc$spcA),
xlab = "Wavelength / nm",
ylab = "Absorbance",
ylim = c(0, 4),
type = "l",
lty = 1,
col = rgb(0.5, 0.5, 0.5, alpha = 0.3)
)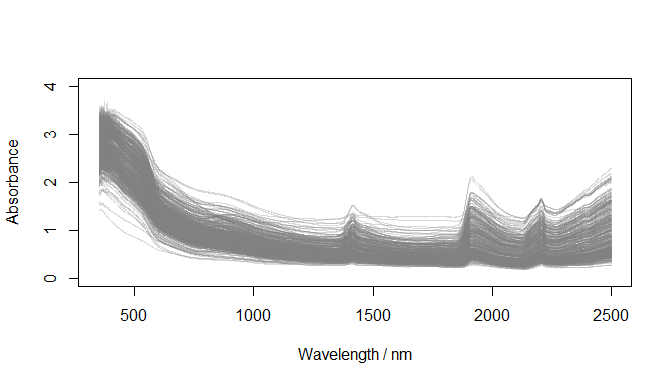
Absorbance spectra (datsoilspc$spcA)
Preprocessing: resample (5 nm) + SNV + moving average
oldWavs <- as.numeric(colnames(datsoilspc$spcA))
newWavs <- seq(min(oldWavs), max(oldWavs), by = 5)
datsoilspc$spcARs <- prospectr::resample(
X = datsoilspc$spcA,
wav = oldWavs,
new.wav = newWavs,
interpol = "linear"
)
datsoilspc$spcASnv <- prospectr::standardNormalVariate(datsoilspc$spcARs)
datsoilspc$spcAMovav <- prospectr::movav(datsoilspc$spcASnv, w = 11)
wavs <- as.numeric(colnames(datsoilspc$spcAMovav))
matplot(
x = wavs,
y = t(datsoilspc$spcAMovav),
xlab = "Wavelength / nm",
ylab = "Absorbance (SNV + movav)",
type = "l",
lty = 1,
col = rgb(0.5, 0.5, 0.5, alpha = 0.3)
)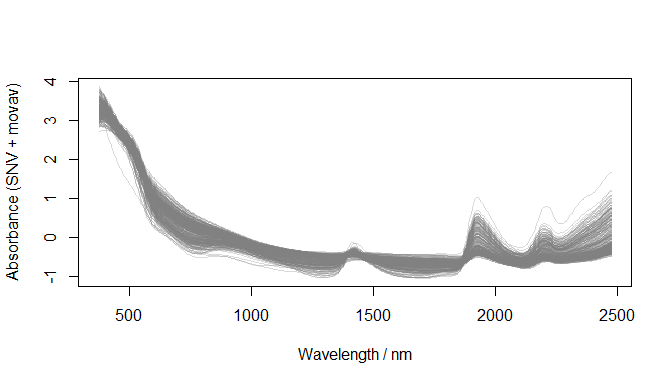
Preprocessed spectra (datsoilspc$spcAMovav)
Split calibration vs validation
set.seed(19101991)
calId <- sample(seq_len(nrow(datsoilspc)), size = round(0.75 * nrow(datsoilspc)))
datC <- datsoilspc[calId, ]
datV <- datsoilspc[-calId, ] # <-- TEST
par(mfrow = c(1, 2))
hist(datC$TotalCarbon, main = "Calibration (datC)", xlab = "Total carbon")
hist(datV$TotalCarbon, main = "TEST (datV)", xlab = "Total carbon")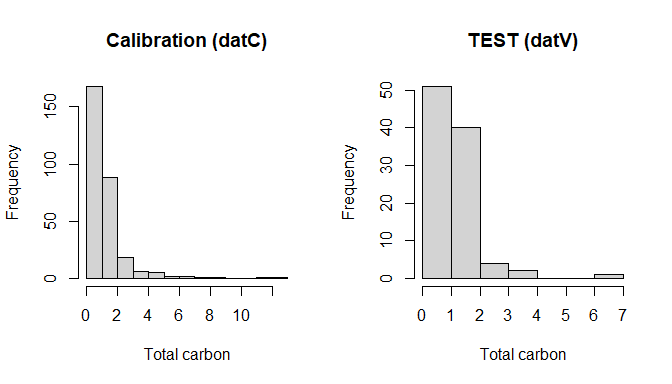
Calibration vs TEST splits (datC vs datV)
Baseline model: PLS
We fit PLS on calibration and evaluate on validation.
maxc <- 30
soilCPlsModel <- pls::plsr(
TotalCarbon ~ spcAMovav,
data = datC,
method = "oscorespls",
ncomp = maxc,
validation = "CV"
)
plot(soilCPlsModel, "val", main = "PLS CV performance (datC)", xlab = "Number of components")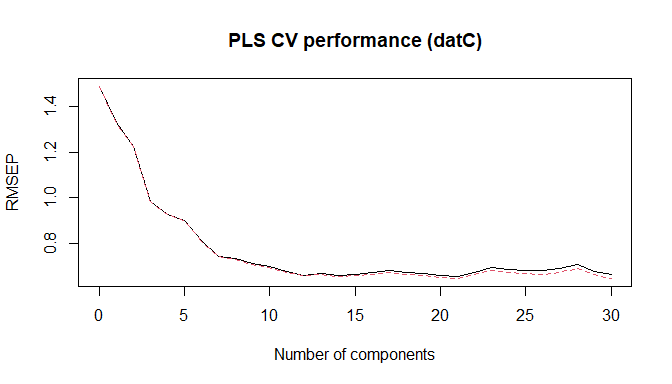
PLS CV performance (datC)
Choose number of components (example uses nc = 14).
nc <- 14
# Refit on full datC with chosen nc (PLS itself uses all comps up to maxc; prediction uses nc)
soilCPlsPred_C <- as.numeric(predict(soilCPlsModel, ncomp = nc, newdata = datC$spcAMovav))
soilCPlsPred_T <- as.numeric(predict(soilCPlsModel, ncomp = nc, newdata = datV$spcAMovav))
par(mfrow = c(1, 2))
plot(datC$TotalCarbon, soilCPlsPred_C, xlab="Observed", ylab="Predicted", main="PLS (datC)", pch=16); abline(0,1)
plot(datV$TotalCarbon, soilCPlsPred_T, xlab="Observed", ylab="Predicted", main="PLS (TEST datV)", pch=16); abline(0,1)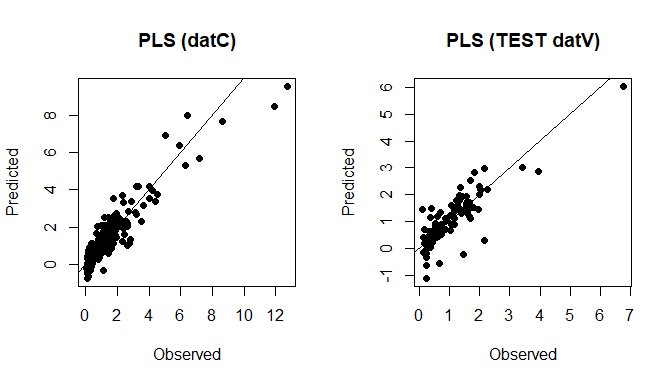
PLS predictions (datC + datV)
Metrics (PLS)
We use the same evaluation function used in many soilspec workflows.
pls_cal <- eval_quant(datC$TotalCarbon, soilCPlsPred_C)
pls_tst <- eval_quant(datV$TotalCarbon, soilCPlsPred_T)
as_df_metrics(pls_cal)
#> n ME MAE RMSE R2 RPIQ RPD
#> 1 293 0 0.37 0.56 0.86 2.04 2.63
as_df_metrics(pls_tst)
#> n ME MAE RMSE R2 RPIQ RPD
#> 1 98 0.02 0.36 0.52 0.69 2.34 1.81Supervised VAE regression: soilVAE
Availability check (TensorFlow/Keras)
This chunk detects if Python + TensorFlow + Keras can be loaded.
If not available, the VAE section is skipped (vignette still builds).
has_py <- reticulate::py_available(initialize = FALSE)
has_tf <- FALSE
if (has_py) {
try(reticulate::py_config(), silent = TRUE)
has_tf <- reticulate::py_module_available("tensorflow") &&
reticulate::py_module_available("keras")
}
has_py
#> [1] TRUE
has_tf
#> [1] TRUEPrepare matrices (same predictors as PLS preprocessing)
Prepare Train/Val internal split within datC (no y transform; scale X only)
set.seed(19101991)
nC <- nrow(datC)
id_tr <- sample(seq_len(nC), size = round(0.80 * nC))
datC_tr <- datC[id_tr, ]
datC_va <- datC[-id_tr, ]
# y stays in original units (no transformation)
y_tr <- as.numeric(datC_tr$TotalCarbon)
y_va <- as.numeric(datC_va$TotalCarbon)
# X: scale predictors using TRAIN center/scale only
X_tr_raw <- as.matrix(datC_tr$spcAMovav)
X_va_raw <- as.matrix(datC_va$spcAMovav)
X_te_raw <- as.matrix(datV$spcAMovav) # TEST
X_tr <- scale(X_tr_raw)
X_center <- attr(X_tr, "scaled:center")
X_scale <- attr(X_tr, "scaled:scale")
# safe scaling: avoid division by zero
X_scale[X_scale == 0] <- 1
X_va <- scale(X_va_raw, center = X_center, scale = X_scale)
X_te <- scale(X_te_raw, center = X_center, scale = X_scale)
# sanity checks (dims)
dim(X_tr)
#> [1] 234 421
length(y_tr)
#> [1] 234
dim(X_va)
#> [1] 59 421
length(y_va)
#> [1] 59
dim(X_te)
#> [1] 98 421Prepare Train/Val internal split within datC (no y transform; scale X only)
We model TotalCarbon using the preprocessed spectra matrix spcAMovav.
Fit + evaluate soilVAE (skipped if TF/Keras unavailable)
reticulate::py_run_string("
import os
import random
import numpy as np
import tensorflow as tf
os.environ['PYTHONHASHSEED'] = '0'
random.seed(19101991)
np.random.seed(19101991)
tf.random.set_seed(19101991)
")
Sys.setenv(TF_DETERMINISTIC_OPS = "1")
Sys.setenv(TF_CPP_MIN_LOG_LEVEL = "2") # reduce logs INFO/WARN
# Sys.setenv(TF_ENABLE_ONEDNN_OPTS = "0")
if (!has_tf) {
message("TensorFlow/Keras not available; skipping soilVAE section.")
} else {
Sys.setenv(TF_ENABLE_ONEDNN_OPTS="0")
# Optional: force a specific python/venv/conda, if needed.
# soilVAE::vae_configure(conda = "soilvae-tf")
grid_vae <- data.frame(
latent_dim = c(8L, 16L, 32L, 64L),
dropout = c(0.20, 0.30),
lr = c(5e-4),
beta_kl = c(0.01),
alpha_y = c(5),
epochs = c(500L),
batch_size = c(64L, 128L),
patience = c(50L),
stringsAsFactors = FALSE
)
grid_vae$hidden_enc <- list(c(512L, 256L, 128L))
grid_vae$hidden_dec <- list(c(128L, 256L, 512L))
tuned <- soilVAE::tune_vae_train_val(
X_tr = X_tr, y_tr = y_tr,
X_va = X_va, y_va = y_va,
seed = 19101991,
grid_vae = grid_vae
)
best <- soilVAE::select_best_from_grid(tuned$tuning_df, selection_metric = "euclid")
cfg <- best$best
# Refit on full datC (train+val) using early stopping monitored on the internal val (datC_va)
m_vae <- soilVAE::vae_build(
input_dim = ncol(X_tr),
hidden_enc = as.integer(strsplit(cfg$hidden_enc_str, "-")[[1]]),
hidden_dec = as.integer(strsplit(cfg$hidden_dec_str, "-")[[1]]),
latent_dim = as.integer(cfg$latent_dim),
dropout = as.numeric(cfg$dropout),
lr = as.numeric(cfg$lr),
beta_kl = as.numeric(cfg$beta_kl),
alpha_y = as.numeric(cfg$alpha_y)
)
soilVAE::vae_fit(
model = m_vae,
X = X_tr, y = y_tr,
X_val = X_va, y_val = y_va,
epochs = as.integer(cfg$epochs),
batch_size = as.integer(cfg$batch_size),
patience = as.integer(cfg$patience),
verbose = 0L
)
yhat_tr <- as.numeric(soilVAE::vae_predict(m_vae, X_tr))
yhat_va <- as.numeric(soilVAE::vae_predict(m_vae, X_va))
yhat_te <- as.numeric(soilVAE::vae_predict(m_vae, X_te))
# Metrics: internal train/val + FINAL TEST
vae_trn <- eval_quant(y_tr, yhat_tr)
vae_val <- eval_quant(y_va, yhat_va)
vae_tst <- eval_quant(as.numeric(datV$TotalCarbon), yhat_te)
# Plots
par(mfrow = c(1, 3))
plot(y_tr, yhat_tr, main="soilVAE (Train)", xlab="Observed", ylab="Predicted", pch=16); abline(0,1)
plot(y_va, yhat_va, main="soilVAE (Val)", xlab="Observed", ylab="Predicted", pch=16); abline(0,1)
plot(as.numeric(datV$TotalCarbon), yhat_te, main="soilVAE (TEST datV)", xlab="Observed", ylab="Predicted", pch=16); abline(0,1)
par(mfrow = c(1, 1))
}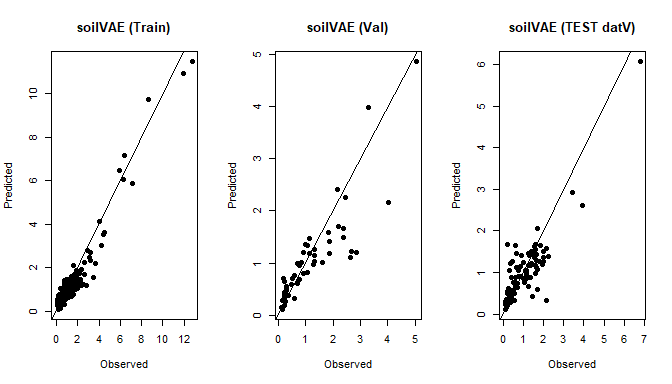
Train/Val splits for soilVAE
Compare PLS vs soilVAE (TEST = datV)
We present a compact comparison table.
if (!has_tf) {
tab <- rbind(
cbind(Model = "PLS", Split = "Calibration (datC)", as_df_metrics(pls_cal)),
cbind(Model = "PLS", Split = "TEST (datV)", as_df_metrics(pls_tst))
)
} else {
tab <- rbind(
cbind(Model = "PLS", Split = "Calibration (datC)", as_df_metrics(pls_cal)),
cbind(Model = "PLS", Split = "TEST (datV)", as_df_metrics(pls_tst)),
cbind(Model = "soilVAE",Split = "Train (internal)", as_df_metrics(vae_trn)),
cbind(Model = "soilVAE",Split = "Val (internal)", as_df_metrics(vae_val)),
cbind(Model = "soilVAE",Split = "TEST (datV)", as_df_metrics(vae_tst))
)
}
row.names(tab) <- NULL
tab
#> Model Split n ME MAE RMSE R2 RPIQ RPD
#> 1 PLS Calibration (datC) 293 0.00 0.37 0.56 0.86 2.04 2.63
#> 2 PLS TEST (datV) 98 0.02 0.36 0.52 0.69 2.34 1.81
#> 3 soilVAE Train (internal) 234 -0.07 0.31 0.44 0.92 2.54 3.60
#> 4 soilVAE Val (internal) 59 -0.10 0.33 0.51 0.76 2.36 2.04
#> 5 soilVAE TEST (datV) 98 -0.04 0.33 0.47 0.74 2.56 1.97If TensorFlow/Keras was not available, you can still use the PLS section and install a compatible Python stack later.
Notes for reproducibility
The PLS workflow depends only on R packages
plsandprospectr.The supervised VAE requires:
Notes for life
Education without ethics is only rhetoric.
Power without reflection is violence


